Research ProjectsSonification of Vital Sign DataNewest project is looking at whether there is a way to represent univariate times series data through music. The work is in the beginning stages and achieved in collaboration with Dr. Carsten Skarke of the Institute of Translational Medicine and Therapeutics at the University of Pennsylvania.The Kavita ProjectThe Kavita Project is focused on developing an integrated development environment (IDE) that uses speech recognition as its primary interface. It is being developed using open source technologies. The project is named after Kavita Krishnaswamy, a PhD student at UMBC who inspired the project. It is designed for anyone who may have limited mobility in their hands, but wants to program. We are now working with Dr. Keith Vertanen and his graduate student, Sadia Nerwin, of the Michigan Technical University. More information can be found here.Multivariate Time Series Analysis of Physiological and Clinical DataThis project is a continuation of my dissertation research. In collaboration with physicians at Johns Hopkins University and my advisors at UMBC, I developed techniques for the Neonatal and Pediatric Intensive Care Units to provide clinical decision support. We created a visualization tool to assist provider's in analyzing the patient's data from a multivariate perspective over time. Clicking on the image will begin a download of a tutorial video of the latest visualization. I am currently working on developing a similarity metric that will assist physicians in determining how similar a patient is to another physiologically and in terms of experiencing similar medical events. I am also expanding the visualization and a presentation on the latest version of the visualization as well as a technique for data mining multivariate time series data can be seen here. Collaborative Research Experience for Undergrads (CREU)The title of this project is "Multivariate Time Series Analysis of Physiological and Clinical Data to Predict Patent Ductus Arteriosus (PDA) in Neonatal Patients." I wrote the grant for this project which is funded by the CRA-W and the CDC (Coalition for Diversity in Computing). I am working with three underrepresented students in computing; one is a computer scientist, another a computer engineer, and yet another a biologist. All three are helping me to complete a survey with residents at Johns Hopkins Hospital to evaluate how well residents can diagnose patients with PDA using our visualization and clinical decision support tool versus the traditional visualizations. More information about the project can be found at http://creu2010-umbc.wikispaces.org.Context Aware Surgical Training (CAST)CAST was developed to create an environment where pervasive technologies, agent technologies, Semantic Web ontologies, logic reasoning, security and privacy policies, and RFID (radio frequency identifier) technology were being developed to extend the capabilities of the MASTRI Center, a surgical training facility located at the University of Maryland Medical School under guidance of Dr. Anupam Joshi and Dr. Tim Finin. The project was in its development stage when I was a part of it, thus my contribution to the project was creating a web interface by which resident's could do training in the MASTRI center on their own time and the application would keep a log of where they were in their training.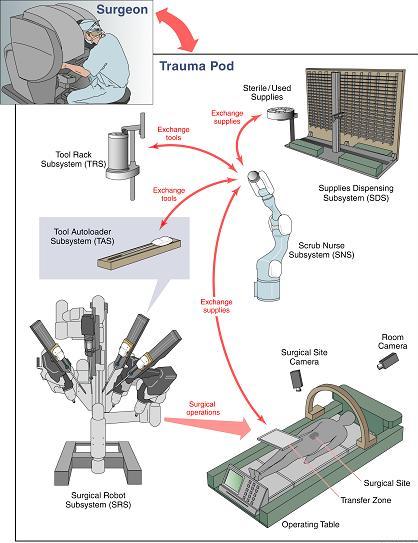 TraumapodIn this project, I worked as part of the eBiquity lab in collaboration with DARPA, SRI, the University of Maryland Medical School, and several other universities to develop a prototype for an unmanned medical treatment vessel that will provide life saving surgical and medical care to soldiers on the battlefield immediately after trauma and during transport. Other partners are developing the robotics that will allow a surgeon to operate on a soldier in the trauma pod remotely. eBiquity's major role in the project was to create and to maintain the Resource Management Subsystem which monitors all the messages that are passed between thesubsystems in order to track the inventory of supplies and tools. We were also responsible for Emergency Medical Record in the project which is an electronic medical record that maintains a time-stamped log of the events in the Trauma Pod for a procedure. My role in the project was to improve the GUI interface, improve the systems capacity to handle the number of messages it receives efficiently, and to serve as the liaison between SRI and UMBC to coordinate and perform the testing. eBiquity's role in the project ended on March 29, 2007. eBiquity's major role in the project was to create and to maintain the Resource Management Subsystem which monitors all the messages that are passed between thesubsystems in order to track the inventory of supplies and tools. We were also responsible for Emergency Medical Record in the project which is an electronic medical record that maintains a time-stamped log of the events in the Trauma Pod for a procedure. My role in the project was to improve the GUI interface, improve the systems capacity to handle the number of messages it receives efficiently, and to serve as the liaison between SRI and UMBC to coordinate and perform the testing. eBiquity's role in the project ended on March 29, 2007.
IBM's Chemical Search EngineWhen I worked in an internship at IBM Almaden Research Center in San Jose, CA in the summer of 2006, I worked with a team of professionals on the IBM Chemical Search Engine. The search engine performs a molecular similarity search for patents and Medline articles and queries for associated metadata by cross referencing the resultant data. My research consisted of creating two interactive visualizations of the resultant data after a query. Previously, the visualization consisted of a single, complex table from which it was difficult to extract information. I constructed two views rather than clutter all the information into a single graph using the interactive visualization toolkit, Prefuse.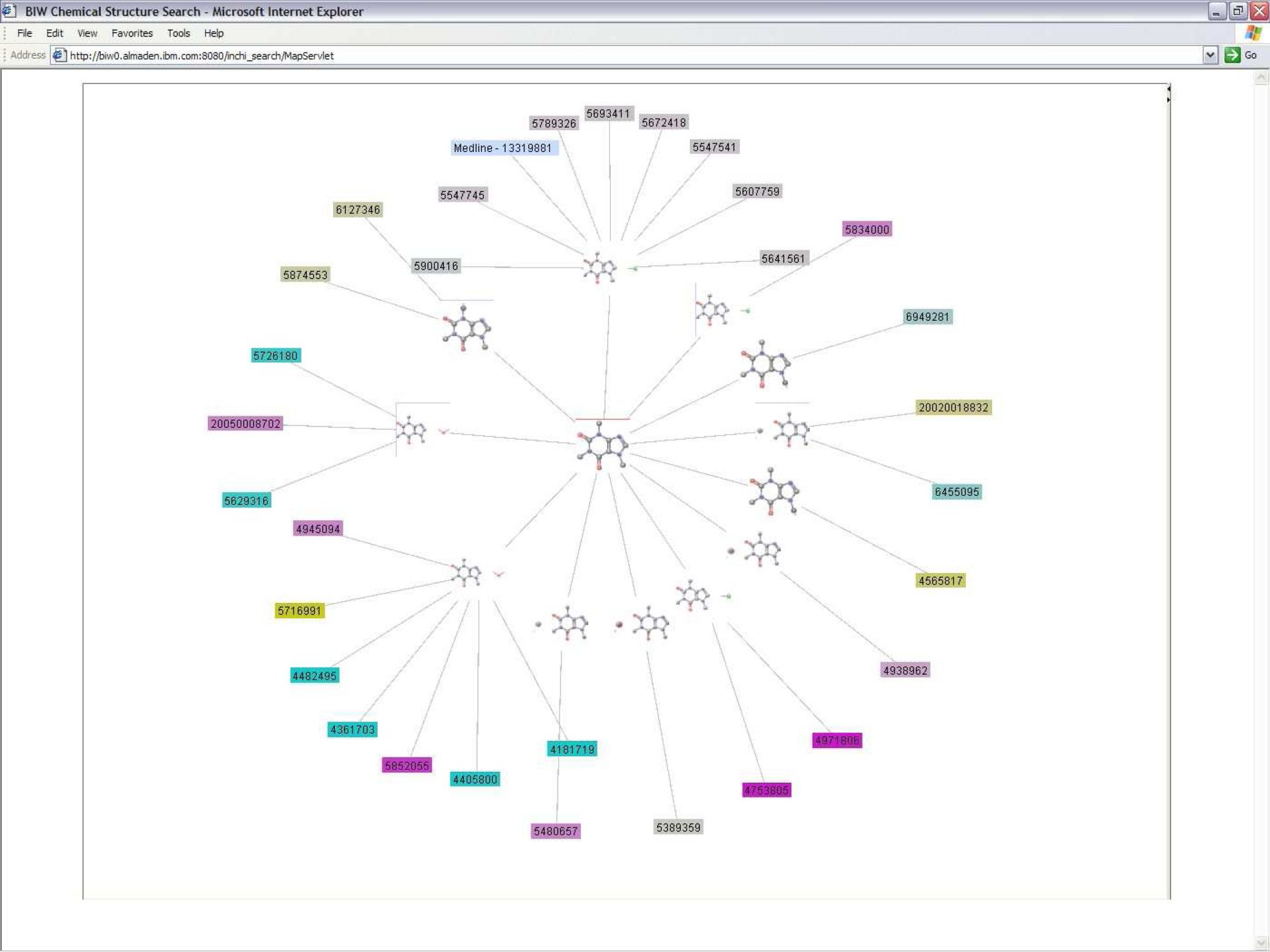 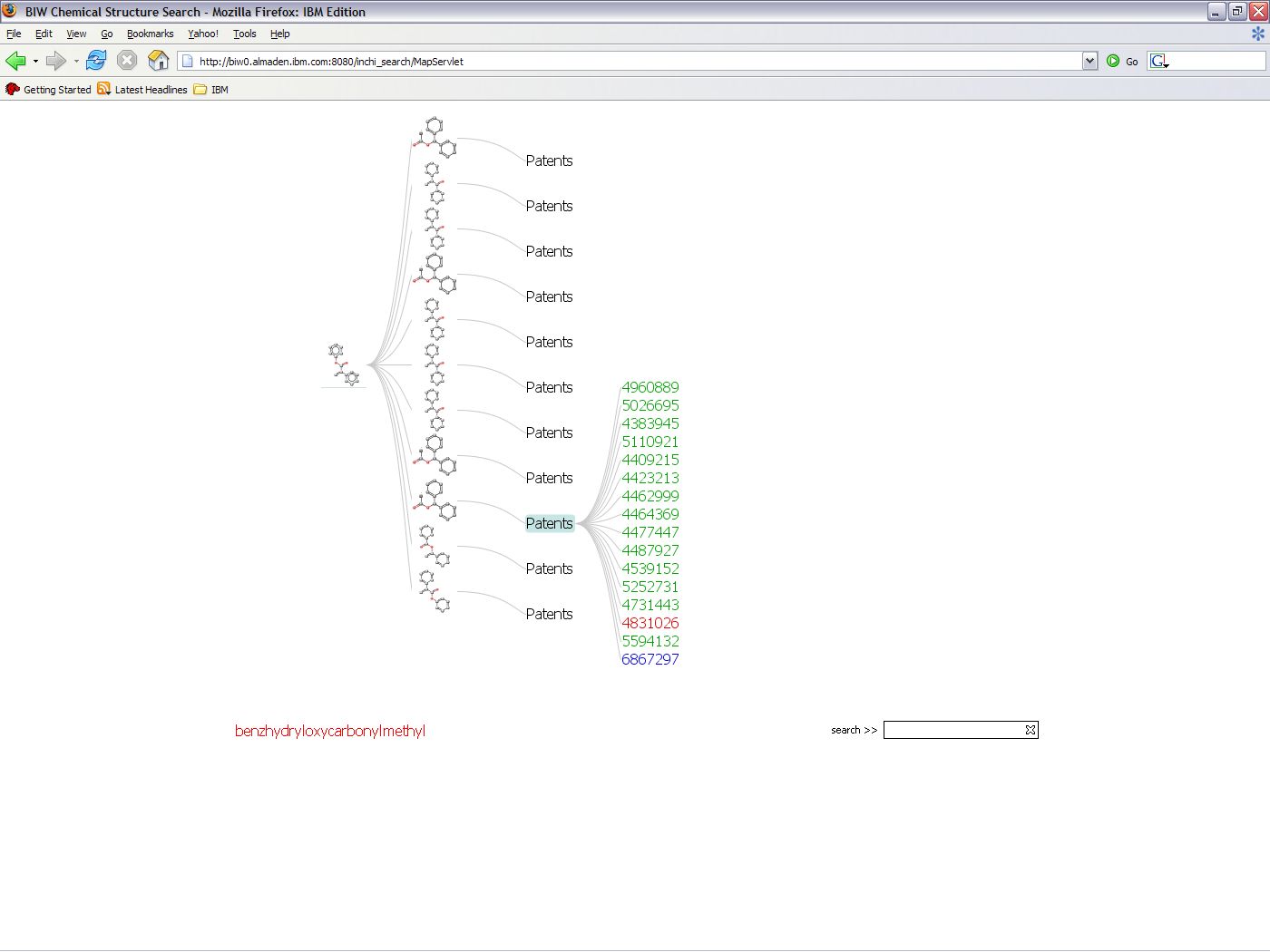 The Graph View, on the left, emphasizes the relationships between patents and compounds, such as which patents name more than one compound. The Tree View, on the right, allows the user to focus on the data for a single compound. A paper on this work was published in the proceedings of the Pacific Symposium on Biocomputing 2002. My contribution to the paper was the generation and analysis of the experimental results. The visualizations were supposed to become a part of the web interface, but as a result of Prefuse and IBM licensing conflicts, it was not possible. Unfortunately, I didn't get a good image of the GraphView before I left. Both visualizations are interactive. If you click on the node with the patent number, the patent is displayed. The Graph View, on the left, emphasizes the relationships between patents and compounds, such as which patents name more than one compound. The Tree View, on the right, allows the user to focus on the data for a single compound. A paper on this work was published in the proceedings of the Pacific Symposium on Biocomputing 2002. My contribution to the paper was the generation and analysis of the experimental results. The visualizations were supposed to become a part of the web interface, but as a result of Prefuse and IBM licensing conflicts, it was not possible. Unfortunately, I didn't get a good image of the GraphView before I left. Both visualizations are interactive. If you click on the node with the patent number, the patent is displayed.
|